PRL3A links regeneration, differentiation and melanoma
Researchers identify pathway that links melanocyte regeneration and cancer: August 2020
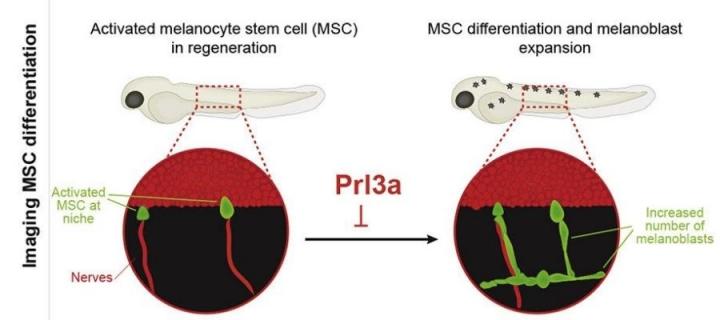
Melanocytes, replenished throughout life by melanocyte stem cells (MSCs), play a critical role in pigmentation and melanoma.
Liz Patton and colleagues reveal a new developmental checkpoint for MSC differentiation in melanocyte regeneration. Unexpectantly, they discover that regulation of transcriptional elongation of endomembrane genes by the metastasis-associated Phosphatase of Regenerating Liver 3 ( PRL3) controls the rate of regeneration of these cells. PRL3 has a well-established function in cancer, and they show here that this new mechanism discovered in MSCs is active in melanomas from patients, suggesting new targets in transcriptional elongation for PRL3-high melanomas.
To understand how melanocytes regenerate from stem cell populations, the researchers screened zebrafish embryos for small molecule compounds that helped melanocytes rapidly regenerate after damage. They discovered one compound, B4-Rhodanine (B4-Rh), and showed that it accelerated the rate of melanocyte regeneration from MSCs.
B4-Rh potently inhibits the Prl3 phosphatase enzyme, so they hypothesised that this inhibition resulted in the increased cell regeneration. Using genetics, they tested this using fish lacking working Prl3 genes and found that if the Prl3a gene was not functional, regeneration was increased and this was not amplified further by B4-Rh. Conversely, when they increased Prl3a in zebrafish embryos this impaired the development of melanocytes suggesting an important role for Prl3a in melanocyte development and regeneration.
Using mass spectrometry and super resolution microscopy methods, the researchers showed Prl3a binds the RNA Helicase Ddx21, an important enzyme for sensing nucleotide pools in the cell, and binding chromatin to regulate elongation of gene transcription. Reducing Ddx21 in zebrafish embryos demonstrated that this was vital for the regeneration response and state-of-the-art microscopy experiments showed Prl3a and Ddx21 colocalised in the nucleus of a melanoma cell line.
Then, in collaboration with Rob Illingworth’s lab (CRM, UoE), they used in vivo metabolic RNA labelling techniques to demonstrate that high levels of PRL3 resulted in less and restricted DDX21 chromatin binding, and shorter RNA transcripts of endolysosomal genes. Remarkably, the same transcriptional pathways were restricted in zebrafish embryos expressing high levels of PRL3. Thus, through cell and biochemical experiments coupled with RNA-seq transcriptomics, the researchers showed that these proteins are important to fine tune the elongation of RNA strands during transcription, and these same pathways were dysregulated in the zebrafish embryos.
Given that PRL3 is a primarily known for its role in cancer, the researchers then went on to show that high PRL3 expression was also significantly associated with reduced expression of endolysosomal genes and melanoma-specific death in two patient melanoma cohorts, suggesting a real relevance to human cancer, most likely as a biomarker for metastatic potential.
With elegant use of cell and zebrafish models, this work presents the conceptual advance that PRL3-mediated control of transcriptional elongation is a differentiation checkpoint mechanism for activated MSCs and has clinical relevance for the activity of PRL3 in regenerating tissue and cancer.
This mechanism controls premature melanoblast expansion and differentiation from MSCs. In melanoma patients, restricted transcription of this endolysosomal vesicle pathway is a hallmark of PRL3-high melanomas.
Links
- Liz Patton Research Group
- 'PRL3-DDX21 Transcriptional Control of Endolysosomal Genes Restricts Melanocyte Stem Cell Differentiation', Developmental Cell: https://doi.org/10.1016/j.devcel.2020.06.013

