Software & Data
Our programs, scripts, and data
Below you can find standalone programs, scripts, and data for reproducing or augmenting our research.
HighlanderLab public repositories of source code and scripts

HighlanderLab code repositories
SIMplyBee - simulating honeybee populations and breeding

Led by a team of bee-enthusiasts ;)
stdpopsim - standard population genetic simulation models

Contributions from a number of lab members
Optimising APY core subsets for scalable genomic prediction

Led by Ivan Pocrnic
AlphaPart - quantifying the drivers of genetic change
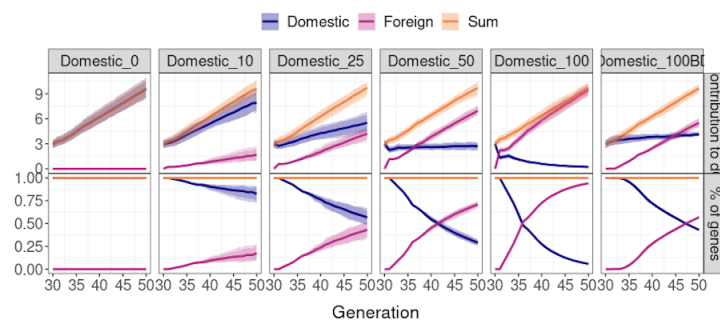
Led by Jana Obsteter and Thiago P. Oliveira
AlphaSimR - flexible modelling of breeding programmes

Led by Chris Gaynor
AlphaMate - optimising selection and management of diversity
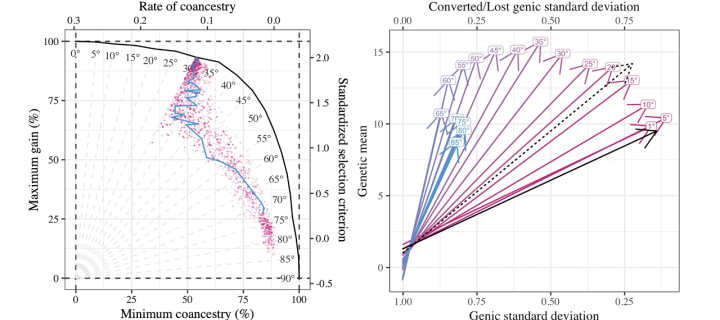
Led by Gregor Gorjanc
AlphaSuite - tools for quantitative genetics, genomics, and breeding
Contributions from a large number of researchers
Temporal and genomic analysis of additive genetic variance
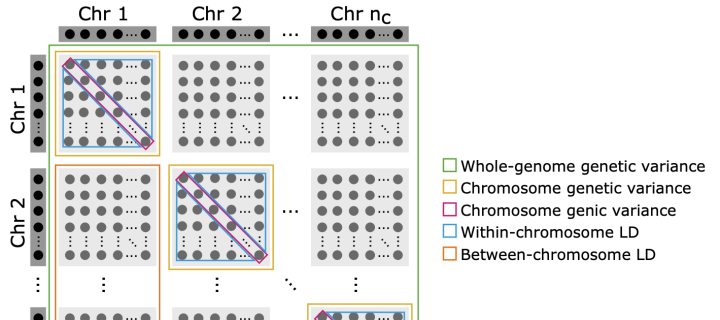
Led by Letícia A. C. Lara
Modelling gene drives to suppress an asian hornet population
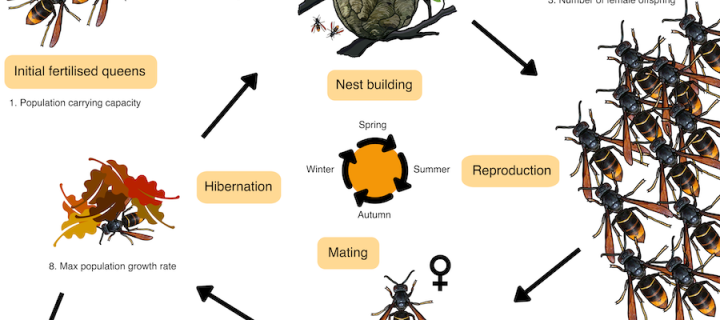
Led by Yani Meiborg
Modelling gene drives to suppress a varroa population
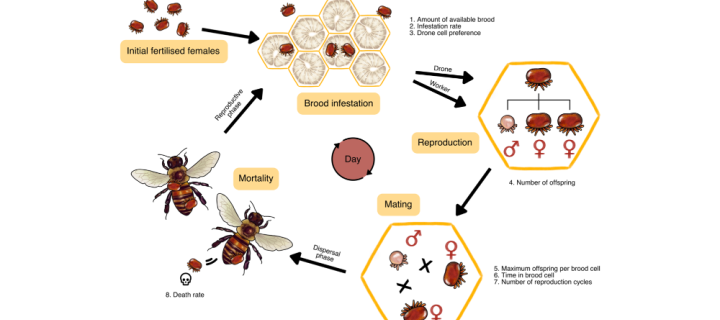
Led by Nicky R. Faber
Modelling gene drives to suppress a grey squirrel population
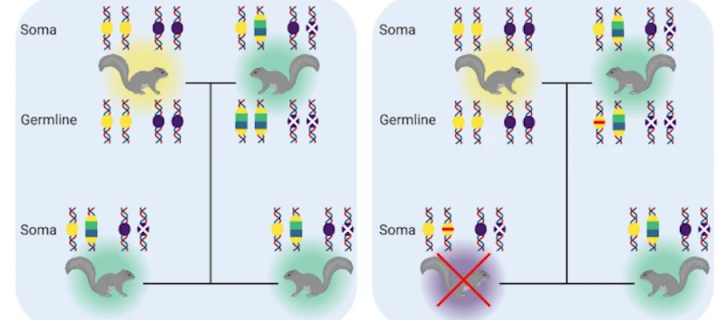
Led by Nicky R. Faber
Modelling gene drives to suppress a mouse population

Led by Nicky R. Faber
Robust modelling of genetic values with expert knowledge
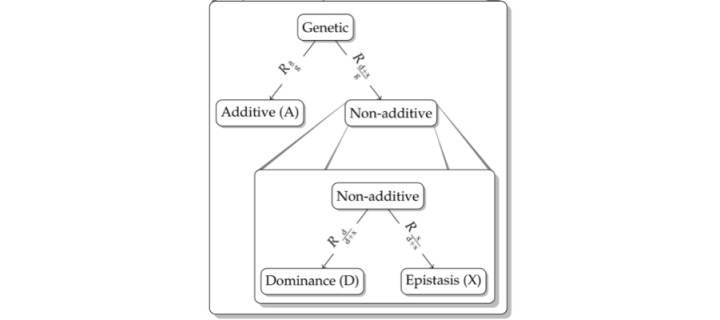
Led by Ingeborg G. Hem
Hierarchical modelling of haplotype effects leveraging tree/phylogeny
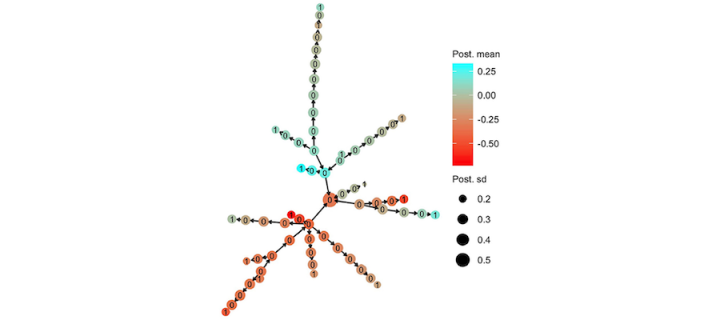
Led by Maria L. Selle
Spatial modelling improves genetic evaluation

Led by Maria L. Selle
Flexible modelling of spatial variation in agricultural field trials
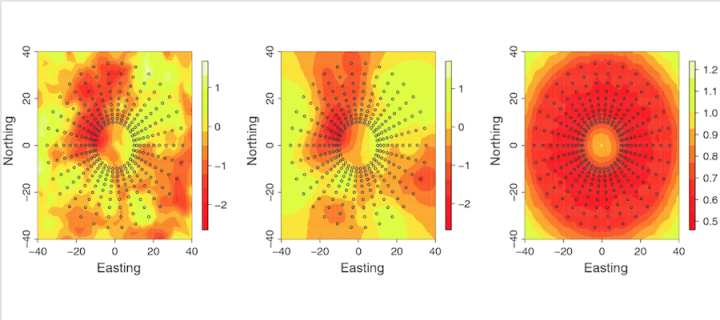
Led by Maria L. Selle

