Genome editing and targeted protein degradation
Our research has three themes:
Theme 1: Degron Tagging
Most strategies to perturb gene function target nucleic acids, whereas proteins are the effectors of most biological functions. Degron tagging enables the dose of specific proteins to be controlled using small molecule ligands that induce turnover via the ubiquitin / proteasome system. Degradation of tagged proteins is often rapid (e.g. t½ < 30 minutes), can be tuned to intermediate/hypomorphic levels, and is reversible once the ligand is removed. This provides greater versatility than many other gene perturbation strategies, making it easier to identify the direct and immediate consequences of protein removal, and allowing protein function to be removed and reinstated to better model gene therapy and drug activity.
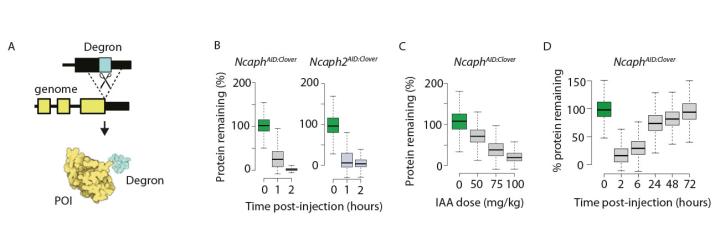
We recently adapted the auxin-inducible degron system for use in mouse models (collaboration with Bin Gu Michigan State University). Endogenously tagged proteins could be degraded effectively in most tissues and cell types, in adults and embryos, and degradation was highly specific for the target protein. However, we found that degradation was more efficient in some cell types compared to others, and for some tagged proteins compared to others, and that different substrates compete for access to a limited ubiquitin ligase pool.
As part of the MRC National Mouse Genetics Network (NMGN) we are developing protocols to solve these problems and broaden the utility of degron tagging across tissues, targets and timescales (https://nmgn.mrc.ukri.org/clusters/degron-tagging/). This is a formidable challenge requiring cross disciplinary collaboration, and we are working with medicinal chemists (Asier Unciti-Broceta lab, CRUK Edinburgh Centre), pharmacologists (Prof. Roland Wolf lab, University of Dundee), protein degradation specialists (Gopal Sapkota lab, MRC PPU) and computational protein biologists (Joe Marsh lab, MRC HGU). What we learn will be translated to models of human disease and therapy through collaborations within the HGU, NMGN and beyond.
Associated Publication:
Lewis Macdonald, Gillian Taylor, Jennifer Brisbane, Ersi Christodoulou, Lucy Scott, Alex Von Kriegsheim, Janet Rossant, Bin Gu*, Andrew J Wood*. Rapid and specific degradation of endogenous proteins in mouse models using auxin-inducible degrons. bioRxiv 2022.01.13.476100; doi: https://doi.org/10.1101/2022.01.13.476100
Theme 2: Genome Editing
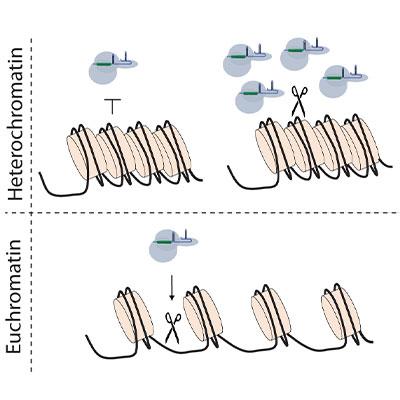
Genome editing enables the sequence of chromosomal DNA to be modified in living cells using engineered nucleases. During his postdoctoral work (UC Berkeley, 2008 – 2011) Andrew led efforts to develop the first protocols for germline genome editing in nematode worms, showing that they could be transferred between species to study the evolution of developmental pathways (Wood, Lo, Zeitler et al, Science, 2011). This work with Zinc Finger and TALE nucleases helped to pave the way for the CRISPR revolution, especially in ‘non-traditional’ model organisms. Since then, the laboratory has helped to pioneer epigenome editing tools in mammalian stem cell models (Therizols et al, Science 2014), and elucidated the contribution of chromatin state to the frequency and type of mutations that arise during CRISPR-Cas9 genome editing (Kallimasioti-Pazi et al, PLoS Biology 2018). Current projects use genome editing technology for deep mutational scanning to better understand genotype/phenotype relationships in human disease genes.
Associated Papers:
- Kallimasioti-Pazi I, Chathoth K, Taylor G, Meynert A, Ballinger T, Kelder M, Sanli I, Lalevee S, Feil R, Wood AJ. Heterochromatin impedes CRISPR-Cas9 mutagenesis but does not influence the outcome of mutagenic DNA repair. PLoS Biology 2018; Dec 12, 16(2):e2005595
- Therizols PT, Illingworth R, Courilleau C, Boyle S, Wood AJ, Bickmore WA. Chromatin decondensation is sufficient to alter nuclear positioning in embryonic stem cells. Science. 2014 Dec 5;346(6214):1238-42.
- *Wood AJ, *Lo TW, *Zeitler B, Pickle CS, Ralston EJ, Lee AH, Amora R, Miller JC, Leung E, Meng X, Zhang L, Rebar EJ, Gregory PD, Urnov FD, Meyer BJ. Targeted genome editing across species using ZFNs and TALENs. Science. 2011 Jul 15;333(6040):307
Theme 3: Mitotic Chromosome Condensation
Successful completion of the cell cycle is essential for multicellular development. However, heritable mutations affecting genes encoding essential cell cycle proteins can cause tissue-specific diseases such as microcephaly, immunodeficiency and cancer predisposition, but the basis for these tissue-specific phenotypes is poorly understood. We discovered (with Bickmore and Jackson) that germline mutations affecting subunits of the condensin complexes, which restructure the genome in preparation for cell division, cause developmental disease through abnormal mitosis (Woodward et al Genes & Development 2016, Martin et al Genes & Development 2016). In mice, thymic T cell divisions are particularly affected by condensin mutation, whereas in human, neurogenesis is disproportionately compromised. We also showed for the first time that perturbation of mitotic chromosome structure was sufficient to drive tumourigenesis in mice (Woodward Genes Dev. 2016 Oct 1;30(19):2173). Somatic mutations in condensin subunits occur at elevated rates in human cancer (Leiserson, Nat Genet. 2015 Feb;47(2):106), so our data support that these rare mutations are bona fide drivers of tumourigenesis.
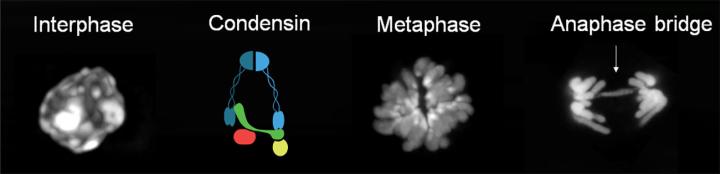
To further investigate the role of condensins during cell divisions in tissues, we generated new CRISPR-engineered mice in which the kleisin subunits of condensin I or condensin II (NCAPH or NCAPH2, respectively) can be rapidly degraded by exposing cells or tissues to indole-3-acetic acid. In a pilot study, we found that early B and T lymphocyte progenitors require both condensin I and condensin II to complete a single round of cell division, whereas either complex can suffice in mature lymphocytes. This provides evidence for specialisation of the mitotic chromosome condensation machinery during tissue development, consistent with our previous findings.
We are using these new mouse lines in our lab and together with collaborators to better understand how condensins regulate chromosome structure during physiological processes within primary cells and tissues.
Associated Publications
- Macdonald L, Taylor G, Brisbane J, Christodoulou E, Scott L, Kriegsheim A, Rossant J, Gu B*, Wood AJ*. Rapid and specific degradation of endogenous proteins in mouse models using auxin-inducible degrons. bioRxiv 2022 doi: https://doi.org/10.1101/2022.01.13.476100
- Woodward J, Taylor GC, 12 others, Arends MJ, Kranc KR, Jackson AP, Bickmore WA*, Wood AJ*. Condensin II mutation causes T cell lymphoma through tissue-specific genome instability. Genes & Development 2016; 30(19) 2173-2186.
- Martin CA, Murray JE, 18 others, Wood AJ, Vagnarelli P, Jackson AP. Mutations in genes encoding condensin complex proteins cause microcephaly through decatenation failure at mitosis. Genes & Development 2016; 30(19):2158-2172
- Wood AJ*, Severson AF*, Meyer BJ. 2010 Condensin and cohesin complexity: the expanding repertoire of functions. Nature Reviews Genetics 11 (6), 391


