Gene insights open doors to scientific advances
Knowledge and understanding of genes and genetics are key to developing healthy, productive animals.
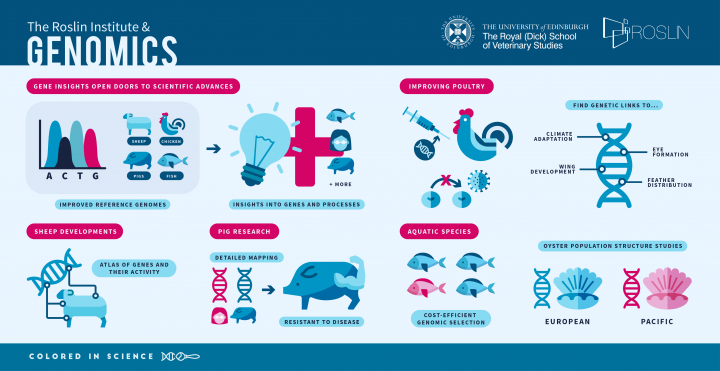
At the Roslin Institute, insight into genes and the processes governing physical and behavioural traits is enabling a wealth of discoveries in biology.
Roslin scientists have developed improved reference data map on the genetic code of key farm animals, including pigs, chickens and sheep as well as key aquatic species.
The improved pig reference genome – which details the whole genetic makeup of pigs – is comparable in detail to the current reference genome for humans.
This mapping effort has supported the production of pigs resistant to a commercially significant disease, porcine reproductive and respiratory syndrome virus, and can be used to target other diseases in pigs in the future.
Ongoing research involves the sequencing of genomes of other pig species including those that differ in resistance to African swine fever – a devastating viral disease threatening global pig production.
Sheep developments
Researchers contributed to a comprehensive atlas of genes and their activity in different parts of the body in domestic sheep. This helps to understand the functions of genes throughout life, enabling extensive research into health, welfare and beneficial breeding.
The availability of such data allowed scientists to develop gene-edited sheep as a model with which to study Batten disease, a devastating degenerative condition in children.
Improving poultry
Researchers have used gene-editing techniques to stop the bird flu virus from spreading in chicken cells grown in the lab.
This was made possible by the methods to culture chicken stem cells and the availability of a high quality genome sequence for domestic chickens.
Improved understanding of the chicken genome has enabled scientists to pinpoint areas of the genome linked to climate adaptation in African poultry.
Further studies in chicken cells and embryos, including with fluorescent markers on key indicator proteins, have allowed scientists to shed light on genes linked to wing development, feather density and distribution, eye formation, and the function of immune cells, demonstrating the possibility of precise cell and tissue studies in embryos.
Aquatic species
Scientists have made significant progress in the development of tools to accelerate genetic improvements in aquaculture.
Researchers have built upon previous work to devise a way to improve the cost-efficiency of genomic selection – a method of choosing the best animals for breeding – in Atlantic salmon. The same approach has been used at Roslin to dissect resistance to amoebic gill disease and inform breeding strategies to produce fish resistant to this disease.
Investigations into the genomes of Pacific and European oysters have defined the structure of European populations and demonstrated the potential for genomic selection of growth traits in Pacific oysters. The data was used to dissect the genetic basis of resistance to viral infection (OsHV-1) and parasite (Bonamia) infestation in oyster populations.
A similar approach was devised to guide breeding of shrimp in Mexico, supported by fellowships from the Mexican National Council of Science and Technology.
Researchers improved genome data for the European sea bass and common carp, and demonstrated how to improve breeding value estimations to accelerate genetic improvement of these species.
Aquaculture research initiative
A strategic partnership has been agreed with WorldFish, including £450,000 funding to develop genomic selection for Nile tilapia.
Work has led to significant funding from the European Union to improve selective breeding of finned fish, as well as £1.1 million for projects funded by BBSRC and NERC.
Further European Union funding has been secured to generate genomic data and functional annotation for six key fish species.
Research will be enabled with the creation of a freshwater aquarium, supported by £700,000 from the Biotechnology and Biological Sciences Research Council and Agri-EPI capital funding.
International partnerships
Roslin scientists have played a driving role in an international initiative to identify functional elements in farm animal genomes. This project, Functional Annotation of Animal Genomes (FAANG), unites international laboratories to inform of understanding of the activity of different parts of the genome and how this can be used to improve animal breeding.
Roslin is a leading contributor to a similar initiative for genomes of fish, including salmon and trout, and the data generated across these projects is freely shared with other researchers and other users.
Roslin researchers are harnessing genome data for hundreds of thousands of animals to inform the design of breeding programmes, a concept termed Genomic Selection 2.0.
This approach to predicting breeding values has attracted industry funding, to support gene analysis of 325,000 pigs with backing from Genus, 289,000 broiler chickens with funding from Aviagen, and more than 1 million cattle with help from the Irish Cattle Breeding Federation.
Research has also identified key parts of the genome linked to Salmonella and Campylobacter resistance in commercial chickens, bovine TB resistance in cattle, resistance to Theileria parasites in African cattle, and resistance of Atlantic salmon to amoebic gill disease and sea lice.
Graphics by Elena Bernabeu at Colored In Science
Related links
Gene-edited pigs are resistant to billion dollar virus
Aquaculture genetics consortium set to tackle industry challenges
Scottish consortiums take giant leap forward for salmon gill health
Study could explain higher rates of human E. coli infection in Scotland
Egg-free surrogate chickens produced in bid to save rare breeds
Briefing on African Swine Fever
The Sheep Gene Expression Atlas
Sheep research could aid insights into childhood dementia


