Liz Patton Research Programme
Research Programme
Melanocyte stem cells in development and human disease
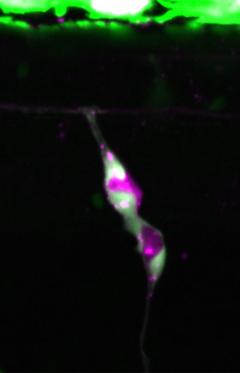
Melanocytes are our pigment cells that protect us from harmful UV-irradiation and colour our hair, skin and eyes. Melanocytes are replenished from reservoirs of melanocyte stem cells (McSCs), although the mechanisms that underpin McSCs for skin melanocytes are not well explored. In zebrafish, skin melanocytes arise directly from the neural crest or via a McSC population that is also neural crest derived and resides at a dorsal root ganglia niche. Little is known about McSCs, however, it is increasingly understood that dysregulated McSC states emerge in melanoma cell populations. Therefore, understanding the genetic control of McSCs may be important for developing new melanoma therapies designed to target this lineage.
In our lab, we use the power of chemical genetics, live imaging, and cellular genomics to identify new mechanisms that control McSCs in zebrafish, and relate these to human disease. Using our zebrafish models, we have recently discovered three new McSC mechanisms. First, we uncovered a new transcriptional elongation checkpoint mechanism that controls differentiation and found this to predict patient outcomes in melanoma. Second, we found that McSCs depend on ALDH2 metabolism to generate the metabolites required for generating melanoblast progeny. While ALDH2 has an important role in protecting the genome from toxic metabolites, our recent finding is important because it is the first demonstration of how ALDH2 regulates metabolism in stem cells for differentiation. Finally, we are excited to find that the transcription factor Tfap2b specifies the McSCs in zebrafish embryos that give rise to multiple cell lineages in the adult animal, including all three pigment cell types and nerve associated cells. Thus, McSCs are specified in embryogenesis, but are adult stem cells with multi-fate potential. Tfap2b is dysregulated in melanoma residual disease, and our findings may point to an unexpected McSC pathway that is dysregulated in human disease.
Now, we are excited to be exploring the genomic landscape of McSCs and how these pathways become dysregulated in disease. We have a long-standing collaboration with Professor Veronica Kinsler (Crick Institute) to understand the how the melanocyte lineage is dysregulated in human mosaic pigmentary syndromes. Recently, led by Dr Hannah Brunsdon (Postdoctoral Fellow), and in collaboration with Professor Robert Semple we have been awarded funding from the CLOVE Syndrome Community and the Chan Zuckerberg Initiative to use the zebrafish as a model for mosaic growth disorders.
Key References:
- Brombin et al., Tfap2b specifies an embryonic melanocyte stem cell population that retains adult multi-fate potential. Cell Reports, 2022. PMID: 35021087
- Brunsdon et al., Aldh2 is a lineage-specific metabolic gatekeeper in melanocyte stem cells. Development 2022. PMID: 35485397
- Johansson et al., PRL3-DDX21 Transcriptional Control of Endolysosomal Genes Restricts Melanocyte Stem Cell Differentiation. Developmental Cell. 2020. PMID:32652076.
- Al-Olabi et al., Mosaic RAS/MAPK variants cause sporadic vascular malformations which respond to targeted therapy. J Clin Invest., 2018. PMID: 29461977.
This work is funded the MRC Human Genetics Unit, Melanoma Research Alliance, and European Research Council (finished 2022).
Targeting Melanoma cellular heterogeneity across disease stages

Melanoma is a highly heterogeneous disease, both between and within cancers. For many patients, initial responses to drug treatments are followed by disease recurrence and drug resistance. Genetic and transcriptional cellular heterogeneity is a major contributor to melanoma drug resistance and disease progression, and we and others have shown that transcriptional cellular heterogeneity can emerge from dysregulation of the melanocyte developmental lineage. Thus, it is likely that combinations of drug targets will be required to improve patient outcomes and to eradicate the disease.
In our lab, we use zebrafish models as our primary discovery tool to discover and target melanoma cell populations. Our zebrafish models are especially powerful for modelling the stages of disease observed in patients with cutaneous melanoma: melanoma growth, regression (due to genetic targeting or our new drug pellet treatment method) and disease recurrence. This enables us to understand the minimal residual disease at the regression site, that includes the stroma and immune cells as well as melanoma persister cells. We have recently used cellular genomics, live-imaging and fate mapping to prove that melanoma persister cells arise from the primary tumour, and that the persister cells directly lead to melanoma recurrent disease. This provides us with new opportunities to use genetic and chemical perturbation experiments to identify subpopulations of persister cells and understand the mechanisms that may be targetable to extend progression free survival.
Once we identify new targets in zebrafish, we use mouse models, human cells, and human melanoma tissues to validate our work with a view towards pre-clinical and clinical studies. One of these targets is ALDH1, a metabolic enzyme that marks cancer stem cell populations, and we are working closely with Professor Asier Unciti-Broceta (CRUK Scotland Centre) to develop new drug-leads to target this cell population. We are delighted to be part of an Melanoma Research Alliance Team Science Award with Richard White (MSKCC), Tim Lionnet (NYU), Itai Yanai (NTU) and Travis Hollman (MSKCC) to target persister cells that drive drug resistance, and have recently been awarded an Melanoma Research Alliance - Rosetree Trust Team Science Award (Opportunity for Women in Science) to work with Len Zon (Boston Children’s Hospital), Megan Insco (Dana Farber/Harvard Cancer Centre), Vijay Sankaran (Boston Children’s Hospital), Jonathan Weissman (MIT) and Richard White (MSKCC) to use cellular barcoding to define melanoma drug resistance and cell of origin.
Key References:
- Travnickova et al., Fate mapping melanoma persister cells through regression and into recurrent disease in adult zebrafish. bioRxiv 2022 (in revision).
- Lu & Patton. Long-term non-invasive drug treatments for adult zebrafish. Disease Models & Mechanisms 2022. PMID: 35394043.
- Travnickova et al., Zebrafish MITF-Low Melanoma Subtype Models Reveal Transcriptional Subclusters and MITF-Independent Residual Disease. Cancer Research, 2019. PMID: 31582381.
- Sarvi et al., ALDH1 bio-activates nifuroxzide to eradicate ALDHHigh melanoma-initiating cells. Cell Chemical Biology, 2018. PMID: 30293938.
-
Patton et al., Melanoma models for the next generation of therapies. Cancer Cell, 2021. PMID: 33545064.
-
Patton et al., Zebrafish disease models in drug discovery: from preclinical modelling to clinical trials. Nat Rev Drug Discov, 2021. PMID: 34117457.
-
Travnickova and Patton. Deciphering Melanoma Cell States and Plasticity with Zebrafish Model. J Invest Dermatol, 2021. PMID: 33340501.
This work is funded the MRC Human Genetics Unit, Melanoma Research Alliance, and European Research Council (finished 2022).
New oral melanoma models
Dr Kelly Blacklock (MRC HGU Research Fellow; Veterinary Surgical Oncologist, The Royal (Dick) School of Veterinary Studies) has recently joined our laboratory to develop a new programme dedicated to understanding the mechanisms of oral mucosal melanoma metastasis. Unlike cutaneous melanoma, oral mucosal melanoma develops in the oral cavities, and is relatively common in dogs. We are currently funded by the Kennel Club Charitable Trust to understand if canine oral mucosal melanoma is similar to human oral melanoma, and to understand if the genes and pathways that altered in disease progression can serve as new drug targets. Our aim with this research is to benefit both canine and human patients with oral melanoma.


