Grzegorz Kudla: RNA Synthetic Biology
RNA Synthetic Biology
Aims
In the Kudla lab, we combine synthetic biology, next-generation sequencing, and computational modelling to study the relationships between RNA sequence, structure, expression, and function.
High-throughput mapping of genotype to phenotype.

As large-scale sequencing projects uncover ever new genetic variants in the human genome, understanding of the consequences of this variation is lagging behind. In our lab, we aim to systematically study the consequences of mutations in selected model genes, by constructing large collections of mutated variants and measuring the phenotypes of the mutants. We pioneered this approach in 2009, when we analyzed the effects of synonymous mutations on gene expression and fitness in Escherichia coli cells. Surprisingly, we found that mRNA folding, but not codon adaptation, is the main determinant of expression. We currently use similar methods to analyse the contribution of synonymous mutations to transcription, splicing, RNA degradation and translation in human cells. Ultimately, we want to learn how to predict the effects of disease mutations on gene expression, and to design genes with optimized expression characteristics.
Although experiments with reporter genes can reveal principles of genotype-phenotype relationship, it is important to validate these principles using endogenous genes. To study the effect of mutations on function in a model gene, we recently synthesized 60,000 random mutants of the yeast small nucleolar RNA gene U3, and we measured the fitness of these mutants in a high-throughput pooled competition assay. Analysis of fitness data revealed the effects of all possible individual mutations, and a network of genetic interactions between pairs of mutations. We currently study the effects of U3 mutations on fitness in a range of environmental and genetic backgrounds. We also use the U3 mutant library to analyse the effects of mutations on RNA-protein binding, and on the abundance of RNA in the cell. In collaboration with other labs, we apply genotype-phenotype mapping approaches to study microRNA biogenesis, to develop RNA sensors of small molecules, and to investigate the role of allostery in protein structure.
RNA Interactions
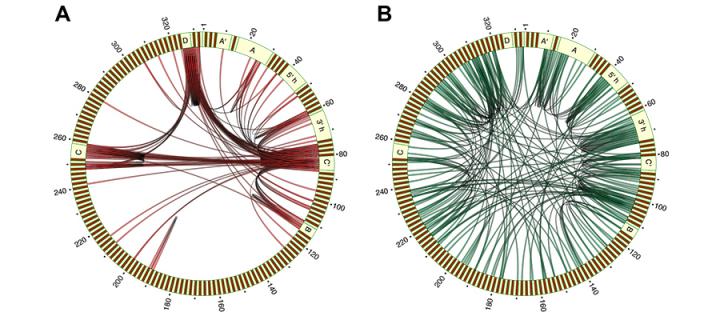
The function of RNA molecules depends on their ability to interact with other molecules, including proteins, RNA, and DNA. We recently described the first high-throughput experimental method for mapping RNA-RNA interactions, called CLASH. CLASH combines affinity purification of protein-RNA-RNA complexes, proximity ligation of interacting RNA molecules, and high-throughput sequencing. We applied this approach for the study of snoRNA-rRNA and microRNA-mRNA interactions, and for mapping the secondary structure of a spliceosomal RNA. We currently use CLASH and related methods for mapping RNA-RNA interactions involved in ribosome biogenesis. We also collaborate with several laboratories to study the role of RNA-protein and RNA-RNA interactions in RNA metabolism, in bacteria, yeast and mammals


