Cohesin is required for long-distance but not proximal enhancer action
Wendy Bickmore's team use protein degradation and synthetic activation approaches to investigate the action of genetic switches called enhancers at various distances: September 2022
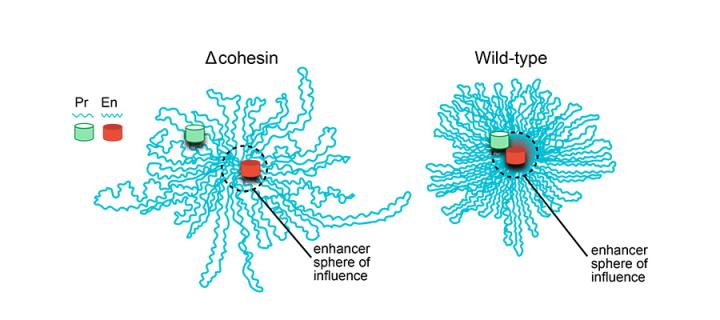
The precise way in which our genes are expressed in development, health and disease is controlled by genetic switches in our DNA called enhancers. Remarkably, enhancers can be found many hundreds of thousands of DNA base pairs away from the target genes that they control. How this long-distance communication occurs is a matter of active debate, but it is generally thought that the three-dimensional organisation of the genome in cells must play a role. By coupling two approaches that allow both 3D genome organisation and gene activation to be precisely controlled experimentally, Kane and colleagues have shown that a molecular machine – termed cohesin - that helps keep the 3D genome compact is required for enhancers to switch on genes located far away in the linear genome. Cohesin is not required for enhancers that are located close to their target gene. These findings help to explain the dynamics of the human genome architecture.
These results gives us a new framework within which to understand how complex gene regulatory landscapes could have evolved to occupy such large regions of the genome, and it puts thinking in 3D centre stage.
Links
Original article: Kane, L., Williamson, I., Flyamer, I.M. et al. Cohesin is required for long-range enhancer action at the Shh locus. Nat Struct Mol Biol (2022). https://doi.org/10.1038/s41594-022-00821-8


