News 2013
An archive of IGMM News from 2013
Islanders to play crucial role in bid to beat killer diseases

Shetland residents are at the forefront of research efforts to pinpoint the causes of heart disease, stroke and diabetes: February 2013
Edinburgh Super-Resolution Imaging Consortium (ESRIC) PhD Studentships
Beginning September 2013 - Two fully-funded 3 year PhD Studentships in high-resolution biological imaging: March 2013
Chancellor's Fellowships

Exciting opportunities for talented postdoctoral scientists at the MRC IGMM: April 2013
Systems Medicine Building

A Brief Description of the Project and New Building: May 2013
IN PICTURES: Inside the heart
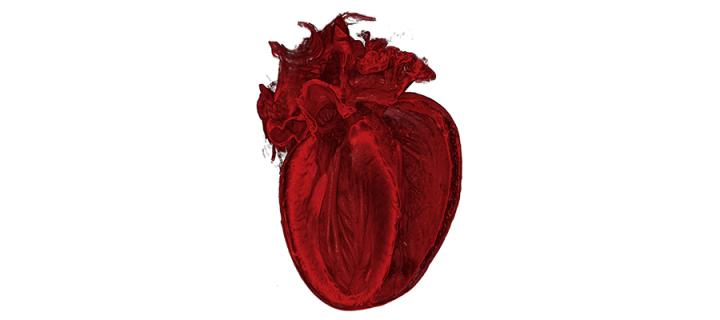
Scientists nationwide submitted videos and images, documenting their research of the heart: July 2013
Athena Swan Bid

IGMM to apply for an Athena SWAN Bronze Award: August 2013
Visit of Her Royal Highness The Princess Royal

Princess Anne visits the Cancer Research UK Edinburgh Centre: October 2013
ECDU and Edinburgh University have sealed a deal with the American drugs Giant Eli Lilly

Edinburgh Cancer Discovery Unit make a deal to develop new drugs to fight cancer: October 2013
3D insight into role of genetics in face development
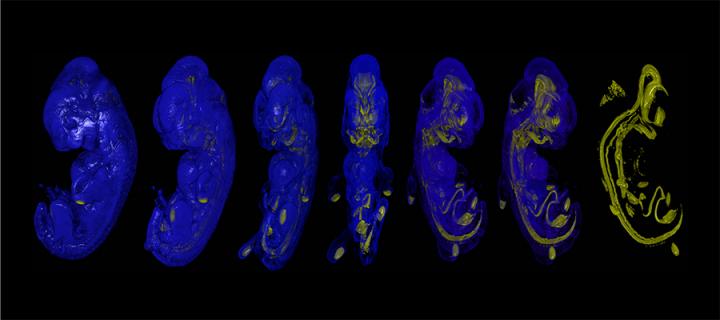
A recent study led by scientists from our MRC Human Genetics Unit, has shown that differences in the activity of gene enhancers drive subtle variation in facial development: November 2013
Promotions and Awards 2013

IGMM researchers recognised for outstanding work: December 2013

